binfalse
Block URLs on phone using AdAway
September 26th, 2015Using AdAway on your Android phones it is very easy to block a set of URLs.
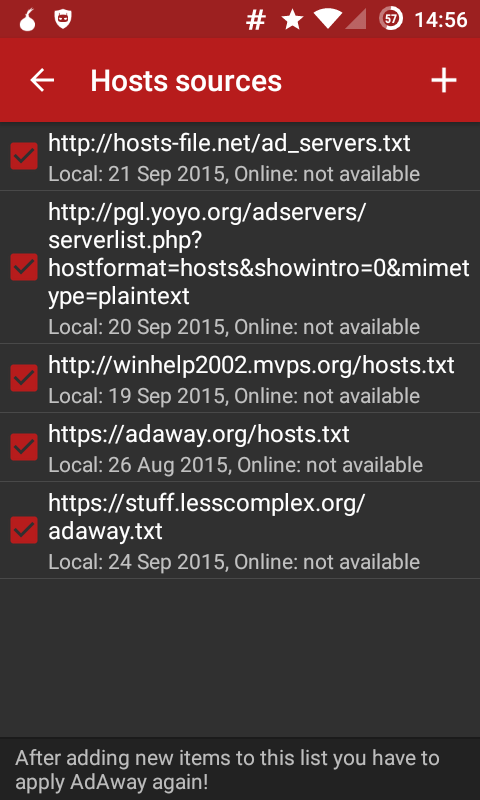
AdAway (available from F-Droid, source from GitHub) is an ad-blocker. The application basically maintains a list of resources containing lists to ad-servers, see figure for example, e.g.:
# dest ip url
127.0.0.1 some.ad.server.com
This entry will redirect all traffic meant for some.ad.server.com to the server 127.0.0.1, which is your localhost. Entries from different resources are merged. Thus, all requests to typical ad-servers will fail → annoying advertisements won’t be delivered. It’s quite powerful and gives impressive results.
However, I didn’t want to talk about ad-stuff. The cool thing is
AdAway allows for extensions with own hosts files!
You can simply add another link to a file containing further host entries. I created my own AdAway file, uploaded it to https://stuff.lesscomplex.org/adaway.txt and added it to AdAway, as you can see in the figure. The current version contains a few entries for:
- facebook/yahoo/etc: I’m not using facebook or stuff, all these request shouldn’t leave my phone
- captive portal detection: To detect captive portals the phone downloads a few bytes from a google server and checks if it is able to access the real internet or just a captive portal
- swift key: I’m using swift key, but do not want it to communicate to the internet. Thus, I’m blocking a few URLs that swiftkey wants to talk to. As I’m already using a firewall this just adds another layer of privacy.
Feel free to use my “extension”, but I expect it to change over time. :)
Shortcomings
This method only works if applications load contents from URLs. As soon as the IPs are hard-coded the hosts file
Disable shutter sound on Cyanogenmod 12
August 26th, 2015Everyone knows that annoying shutter sound of the camera app on Android phones. It’s against the law to sell android phones which do not make sounds when taking pictures. And in general, it is a good feature as it improves other people privacies.
However, I still want to get rid of the sound. It’s a bit tricky, but having a rooted phone (with e.g. CyanogenMod) it’s very easy: Just delete the sound file /system/media/audio/ui/camera_click.ogg! :)
For the lazy: Get a root shell (eg. ssh or adb) and execute the following:
mount -o remount,rw /system
mv /system/media/audio/ui/camera_click.ogg /system/media/audio/ui/camera_click.ogg.backup
mv /system/media/audio/ui/camera_focus.ogg /system/media/audio/ui/camera_focus.ogg.backup
mount -o remount,ro /systemProblem solved.
Kile menu bar hides entries
May 27th, 2015Kile, KDE’s Integrated LaTeX Environment, has a weird bug: Every time I update something in the UI it recreates its config file and I loose some menu options, such as Settings -> Configure Kile where you used to configure your preferences..
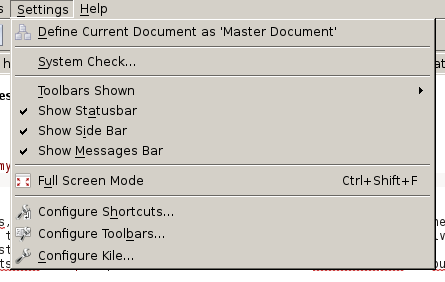
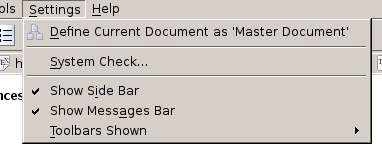
In Figure 1 you can see the menu as expected. There are some entries to Configure Kile, t0 Configure Toolbars, to Configure Shortcuts and to switch to Full Screen Mode, etc. However, as soon as I update certain things in the user interface (UI), eg. if I add a new action icon to the toolbar to quickly get the \textbf{} environment for bold fonts, these entries get lost. In those cases Kile won’t be configureable anymore. The resulting toolbar is shown in Figure 2. You see, the number of entries significantly decreased..
However, I just discovered the reason: Kile in these cases rewrites its config in an unexpected manner…
If you have a look at the configuration stored in ~/.kde/share/apps/kile/kileui.rc you’ll find an XML subtree such as:
...
<Menu noMerge="1" name="settings">
<text>&Settings</text>
<Action name="Mode"/>
<Separator/>
<Action name="settings_perform_check"/>
<Separator/>
<Action append="show_merge" name="StructureView"/>
<Action append="show_merge" name="MessageView"/>
</Menu>
...No idea where it comes from, but the 1 in noMerge prevents the default menu entries to be merged into the minimal set of entries defined in that snippet. However, now that we know what’s wrong we can easily fix it! Just replace the 1 with a 0 and restart Kile. You might want to do the same for all other menus to discover that you’ve also been missing some entries in the Help menu ;-)
Food equivalents in Social Media
May 21st, 2015I’ve just been listening to a podcast from DRadio Wissen: Hörsaal (German podcast streaming lectures). I particularly like the podcast of DRadio. This time the lecture was held by Fabian Hemmert, a German design researcher, who was talking about the smartphones’ future. Quite interesting thought in there, but what I actually wanted to jot down was his comparison between food and social media. He tried to find food equivalents in media:
The equivalent of Fat is Fiction!
Invented/fictional things – easy to consume, but not a good base. And too much of it and you’ll often have unnecessary ballast at the end of the day.
The equivalent of Sugar is Approbation!
Everyone likes to hear approvals and confirmation, it’s music to your ears. But too much of it is also unhealthy.
The equivalent of Protein is Truth!
Not too easy to digest, but an important thing to base on.
We are very short on Attention
He also notes that attention is the scarcest resource. Everyone wants us to pay attention and we want everyone to pay attention..
I kind of like the idea! Any other equivalences you can think of?
Tunneling TinyTinyRSS traffic through a Proxy
May 6th, 2015TinyTinyRSS (TT-RSS) is something that the Google reader always wanted and Feedly still wants to be. Just better! :)
TT-RSS is a free and open source feed aggregator, which can be deployed to your own machine. For example, my instance is running on a cubieboard in my living room. Thus, I’m independent of any company and their plans with my data :)
However, I don’t want to advertise TT-RSS too much, but I want to tell you how to fetch your feeds through a proxy, such as polipo or squid.
Configuring TT-RSS to use a Proxy
It’s apparently undocumented, but looking into the code it turns out that feeds are fetched using cURL:
$ grep -rn PROXY *
include/functions2.php:2257: if (defined('_CURL_HTTP_PROXY')) {
include/functions2.php:2258: curl_setopt($curl, CURLOPT_PROXY, _CURL_HTTP_PROXY);
include/functions.php:389: if (defined('_CURL_HTTP_PROXY')) {
include/functions.php:390: curl_setopt($ch, CURLOPT_PROXY, _CURL_HTTP_PROXY);
plugins/af_unburn/init.php:41: if (defined('_CURL_HTTP_PROXY')) {
plugins/af_unburn/init.php:42: curl_setopt($ch, CURLOPT_PROXY, _CURL_HTTP_PROXY);And as you can see, the code already supports the usage of a proxy: if (defined('_CURL_HTTP_PROXY')).
I think that might be very interesting to many of you guys and I’ve no idea why it is not documented. However, you can simply define the variable _CURL_HTTP_PROXY in your config.php file. For example, to use a proxy at host 127.0.0.1 listening at port 8123 add the following:
define ('_CURL_HTTP_PROXY', '127.0.0.1:8123');Now, the TT-RSS traffic will go through the proxy at :8123, which might tunnel everything through, e.g., TOR. Thus, the location of your living room will not be disclosed :)
BONUS: Cache all the Images in Feeds
By default, TT-RSS will not cache the images in feeds. That means, if there is an image in an article, you will be redirected to load the image from a foreign server. That’s obviously something I’d like to avoid, especially because there are plenty of ads or tracking pixels which shouldn’t know about my habits and surf times. But there is an alternative: TT-RSS is able to cache images. It will download the images to your server and deliver the cached versions instead of forwarding you to somewhere else.
Unfortunately, that is not the default. If you want that functionality you need to configure every single feed (Edit Feed → Options → Cache images locally). And you must not forget to repeat that procedure for every new feed that will be added in 15+ months…
To avoid that you can simply open the database that TT-RSS uses (e.g. using phpMyAdmin), go to the table ttrss_feeds and modify the default value of the column cache_images from 0 to 1. If that is done, the images of every newly added feed will be cached by default.
If you’re too lazy to manually update the feeds that are already there you can simply run the following SQL query:
UPDATE `ttrss_feeds` SET `cache_images`=1 WHERE 1