binfalse
Do I have a CD-RW?
January 6th, 2016You don’t know whether the CD drive on your machine is able to burn CDs? And too lazy to go off with your head under your table? Or you’re remote on the machine? Then that’s your command line:
$ cat /proc/sys/dev/cdrom/info
CD-ROM information, Id: cdrom.c 3.20 2003/12/17
drive name: sr0
drive speed: 32
drive # of slots: 1
Can close tray: 1
Can open tray: 1
Can lock tray: 1
Can change speed: 1
Can select disk: 0
Can read multisession: 1
Can read MCN: 1
Reports media changed: 1
Can play audio: 1
Can write CD-R: 1
Can write CD-RW: 1
Can read DVD: 1
Can write DVD-R: 1
Can write DVD-RAM: 1
Can read MRW: 1
Can write MRW: 1
Can write RAM: 1
Docker Jail for Skype
January 4th, 2016As I’m now permanently installed at our University (yeah!) I probably need to use skype more often than desired. However, I still try to avoid proprietary software, and skype is the worst of all. Skype is an
obfuscated malicious binary blob with network capabilities
as jvoisin beautifully put into words. I came in contact with skype multiple times and it was always a mess. Ok, but what are the options if I need skype? So far I’ve been using a virtual box if I needed to call somebody who insisted on using skype, but now that I’ll be using skype more often I need an alternative to running a second OS on my machine. My friend Tom meant to make a joke about using Docker and … TA-DAH! … Turns out it’s actually possible to jail a usable skype inside a Docker container! Guided by jvoisin’s article Running Skype in docker I created my own setup:
The Dockerfile
The Dockerfile is available from the skype-on-docker project on GitHub. Just clone the project and change into the directory:
$ git clone https://github.com/binfalse/skype-on-docker.git
$ cd skype-on-docker
$ ls -l
total 12
-rw-r--r-- 1 martin martin 32 Jan 4 17:26 authorized_keys
-rw-r--r-- 1 martin martin 1144 Jan 4 17:26 Dockerfile
-rw-r--r-- 1 martin martin 729 Jan 4 17:26 README.md
The Docker image is based on a Debian:stable. It will install an OpenSSH server (it exposes 22) and download the skype binaries. It will also install the authorized_keys file in the home directories of root and the unprivileged user. Thus, to be able to connect to the container you need to copy your public SSH key into that file:
$ cat ~/.ssh/id_rsa.pub >> authorized_keys
Good so far? Ok, then go for it! Build a docker image:
$ docker build -t binfalse/skype .
This might take a while. Docker will execute the commands given in the Dockerfile and create a new Docker image with the name binfalse/skype. Feel free to choose a different name..
As soon as that’s finished you can instantiate and run a new container using:
$ docker run -d -p 127.0.0.1:55757:22 --name skype_container binfalse/skype
This will start the container as a daemon (-d) with the name skype_container (--name skype_container) and the host’s port 55757 mapped to the container’s port 22 (-p 127.0.0.1:55757:22).
Give it a millisecond to come up and then you should be able to connect to that container via ssh. From that shell you should be able to start an configure skype:
$ ssh -X -p 55555 docker@127.0.0.1
The programs included with the Debian GNU/Linux system are free software;
the exact distribution terms for each program are described in the
individual files in /usr/share/doc/*/copyright.
Debian GNU/Linux comes with ABSOLUTELY NO WARRANTY, to the extent
permitted by applicable law.
Last login: Mon Jan 4 23:07:37 2016 from 172.17.42.1
$ skype
You can immediately go and do your chats and stuff, but you can also just configure skype. Do setup everything just like you want to find it when starting skype, for example tick the auto-login button to get rid of the login screen etc. As soon as that’s done, commit the changes to build a new image reflecting your preferences:
$ docker commit skype_container binfalse/deb-skype
Now you’ll have an image called binfalse/deb-skype that contains a fully configured skype installation. Just kill the other container:
$ docker stop skype_container
$ docker rm skype_container
And now your typical workflow might look like:
docker run -d -p 127.0.0.1:55757:22 --name skype__ binfalse/deb-skype
sleep 1
ssh -X -p 55757 docker@127.0.0.1 skype && docker rm -f skype__
Feel free to cast it in a mould just as I did.
The script is also available from my apt repo, it’s name is bf-skype-on-docker:
echo "deb http://apt.binfalse.de binfalse main" > /etc/apt/sources.list.d/binfalse.list
apt-get update && apt-get install bf-skype-on-docker
Getting into a new group
December 30th, 2015You know, … you just got this new floppy disk with very important material but you cannot access it because you’re not in the system’s floppy group and, thus, you’re not allowed to access the floppy device. Solution is easy: add your current user to the floppy group! Sounds easy, doesn’t it? The annoying thing is that those changes won’t take affect in the current session. You need to log out and log in again – quite annoying, especially if you’re into something with lots of windows and stuff. Just happened to me with docker again..
However, there are two methods to get into the new groups without the need to kill the current session:
- su yourself: let’s say your username is
mynameyou just need tosu mynameto get a prompt with the new group memberships. - ssh localhost: that also gives you a new session with updated affiliations.
That way, you do not need to start a new session. However, you still need to start all applications/tools from that terminal - might be odd to those who are used to the gnome/kde menues.. :)
Supplemental material
Display group membership:
groups USERNAME
Add a new system group:
groupadd GROUPNAME
Add a user to a group:
useradd -G GROUPNAME USERNAME
CyanogenMod Updates and the firewall
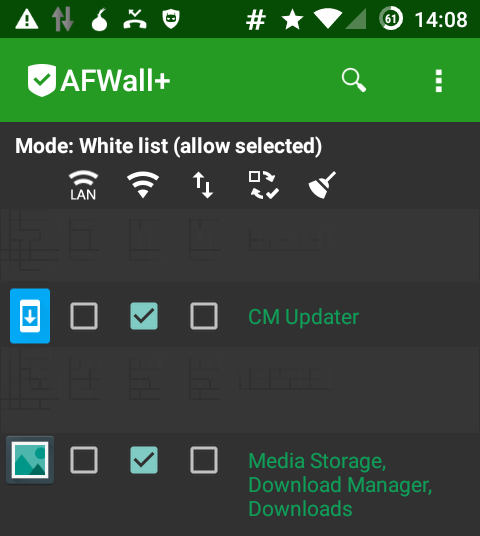
November 3rd, 2015I’m running CyanogenMod on my phone and I have the firewall AFWall+ installed.
To update the list of available updates the application CM Updater need to be able to connect to the internet. It will contact a website from within the google empire and ask for available images.
In order to download a new image the application Media Storage, Download Manager, Downloads needs internet access, as this app is in charge of downloading the actual image file.
To be remembered.
JavaDoc Cheat Sheet
October 5th, 2015- The first sentence of each doc comment should be a summary sentence
- Write the description to be implementation-independent, but specifying such dependencies where necessary. (what is required, what is allowed)
- wrap keywords in
<code>...</code>
typical meta annotations
@authoris not critical, because it is not included when generating the API specification@versionSCCS string"%I%, %G%", which converts to something like “ 1.39, 02/28/97” (mm/dd/yy) when the file is checked out of SCCS@sincespecify the product version when the Java name was added to the API specification (if different from the implementation)
typical method definition
@param parameter-name description- followed by the name (not data type) of the parameter, followed by a description of the parameter
- the first noun in the description is the data type
@return description- omit for methods that return void and for constructors
- include it for all other methods, even if its content is entirely redundant with the method description
@throws class-name descriptionshould be included for any checked exceptions (previously, it was@exception)
additional annotations
{@value package.class#field}- when
{@value}is used (without any argument) in the doc comment of a static field, it displays the value of that constant:The value of this constant is {@value}. - otherwise it displays the value of the specified constant:
Evaluates the script starting with {@value #SCRIPT_START}.
- when
{@code text}- Equivalent to
<code>{@literal}</code>.
- Equivalent to
linking
@see reference- Adds a
"See Also"heading referenceis astring: Adds a text entry for string. No link is generated. The string is a book or other reference to information not available by URL.referenceis an<a href="URL#value">label</a>: Adds a link as defined by URL#value. TheURL#valueis a relative or absolute URL.referenceis anpackage.class#member label: Adds a link (with optional visible text label) that points to the documentation for the specified name in the Java Language that is referenced.
- Adds a
{@link package.class#member label}- in-line link with visible text label that points to the documentation for the specified package, class or member name of a referenced class
- not necessary to add links for all API names in a doc comment
@serial(or@serialFieldor@serialData) for interoperability with alternative implementations of a Serializable class and to document class evolution
deprecation
@deprecated deprecated-text- the first sentence should at least tell the user when the API was deprecated and what to use as a replacement
- a
{@link}tag should be included that points to the replacement method
example
/**
* The Class CombineArchive to create/read/manipulate/store etc.
* CombineArchives.
* <p>
* We directly operate on the ZIP file, which will be kept open. Therefore, do
* not forget to finally close the CombineArchive when you're finished.
*
* @see <a href="https://sems.uni-rostock.de/projects/combinearchive/">
* sems.uni-rostock.de/projects/combinearchive</a>
* @author martin scharm
*/
public class CombineArchive
extends MetaDataHolder
implements Closeable
{
/* ... */
/**
* Gets the the first main entry of this archive, if defined. As of RC2 of the spec there may be more than one main entry, so you should use {@link #getMainEntries()} instead.
*
* @return the first main entry, or <code>null</code> if there is no main entry
* @deprecated as of version 0.8.2, replaced by {@link #getMainEntries()}
*/
public ArchiveEntry getMainEntry ()
{
if (mainEntries == null)
return null;
return mainEntries.size () > 0 ? mainEntries.get (0) : null;
}
/**
* Gets the main entries as defined in the archive.
*
* @return the main entries in this archive
*/
public List<ArchiveEntry> getMainEntries ()
{
return mainEntries;
}
/**
* Replace the file associated with a certain entry while keeping the meta
* data.
*
* @param toInsert
* the new file to insert
* @param oldEntry
* the old entry whose file should be replaced
* @return the new entry
* @throws IOException If an input or output exception occurred
*/
public ArchiveEntry replaceFile (File toInsert, ArchiveEntry oldEntry) throws IOException
{
addEntry (toInsert, oldEntry.getFilePath (), oldEntry.getFormat (), false);
entries.put (oldEntry.getFilePath (), oldEntry);
return oldEntry;
}
}